
The first International Workshop on
Health Natural Language Processing
(HealthNLP 2018)
Organized in conjunction with IEEE ICHI 2018
June 7th, 2018, New York City, USA
Introduction
Natural Language Processing (NLP) technologies have received significant attention in the medical domain and have demonstrated numerous successful uses in healthcare applications. The 2018 HealthNLP workshop will provide a unique platform for close interactions among students, scholars, and industry professionals who are interested in NLP over health data such as clinical notes, social media, and biomedical literature. This new workshop will be held in conjunction with the IEEE International Conference on Healthcare Informatics in New York City on June 7th, 2018.
HealthNLP 2018 invites papers and abstracts about original research and works in progress in natural language processing over health data, focusing on methods, tools, and applications. All submitted papers and abstracts will be peer-reviewed, and selected papers and abstracts will be featured as oral / poster presentations at the workshop. The day-long workshop will also feature a keynote speech and a roundtable discussion.
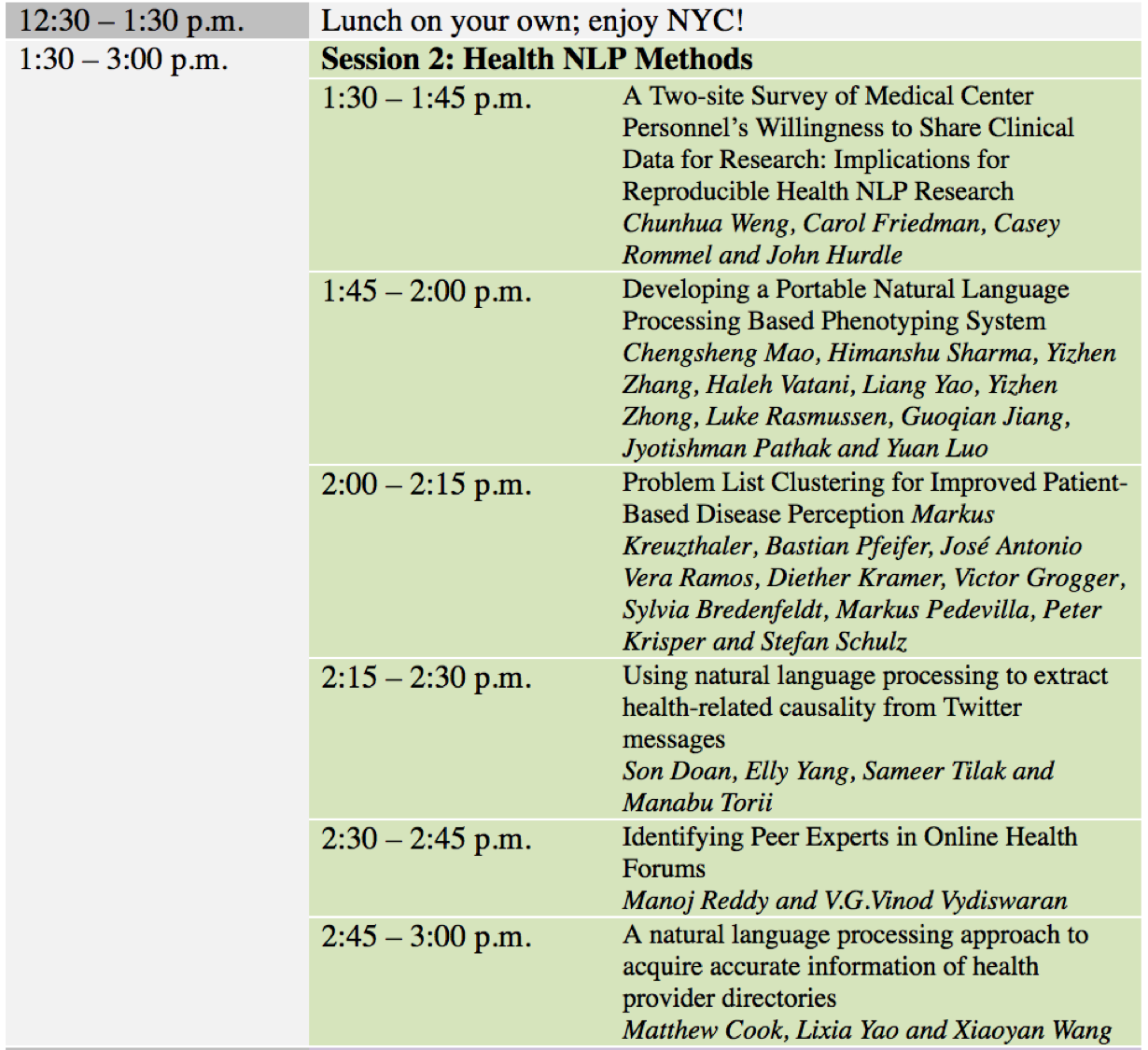
Agenda


Best Paper Award
Thanks to the support from Med Data Quest (MDQ), San Diego, California, we will announce an award of the best paper of HealthNLP 2018 at the workshop. The winner will receive a cash award of $500.
Keynote speaker
Professor Jon Patrick is the co-founder and CEO of HLA, which later become HLA-Global, in 2012. Jon has held many academic positions in his career, most recently the Chair of Information Systems at Sydney University from 1998-2002, and then transferred to the Chair of Language Technology from 2002-2012. In 2012, he left the University to develop his research ideas at HLA; although he still holds an Honorary appointment at Sydney.
In 2005, he won the national Australian Computer Society Eureka Prize for ICT Innovation for his Scamseek project, a system built for the Australian Securities and Investment Commission (ASIC) to identify financial scams on the Internet. Since then, he has applied his expertise to language used in clinical documentation and demonstrated many innovations in advanced processing of clinical documents. In 2010, his team won the international i2b2 Challenge for clinical language processing and has ranked amongst the top teams since.
Title: Language Engineering: Transitions from Research to Industrial NLP
Abstract:
The field of Clinical NLP (CNLP) is dominated by research groups in various hospital and university settings producing investigations for publication or processing systems for local usage. The processing power of these systems are characterised to various levels of complexity by various combinations of small sample sizes, small processing objectives, single source of data, and single purpose utilisation. These systems do not migrate well to larger scale problems where the variety of authors, collection systems, content, and processing objectives are much greater.
Language Engineering (LE) is defined as the process of moving Statistical NLP as performed in the research setting into a context where the scale of complexity is much larger on all the various variables. The core objective is to achieve as high an accuracy as the client demands for Clinical Concept Recognition, which is then used for serving the true user objectives, such as clinical coding, document workflow management, case identification, etc.
An essential process for LE is to develop highly efficient methods for creating the operational system. This requires rapid development of training materials, efficient evaluation of trained language models against the training materials, domain knowledge to direct algorithm development for the pre-processing NLP pipelines and the post-processing which the Clinical Concept Recogniser is intended to serve. These requirements drive a different perspective on the essential requirements for an SNLP pipeline where standard assessment such as 10-fold cross validation has little practical value, where attention to minutiae of processing is vital to achieving project accuracy requirements, and keeping the client properly informed is essential to maintaining their engagement in the project. These principles will be elucidated in a case study for building an automated pathology report coding system for the California Cancer Registry. The beginnings of a workbench to embody these principles have been established in the CLEW project (Clinical Language Engineering Workbench) being developed by the CDC.
Topic of interest
The topics of interest for the papers and abstracts include:
NLP methods: Original methodological research, including but not limited to the following areas: named entity recognition, word sense disambiguation, relation extraction, syntactic parsing, semantic role labeling, topic modeling, discourse analysis, and deep learning
NLP software tools: Any general or specific NLP tools for health-related text data, including clinical notes, biomedical literature, and social media.
NLP applications: Novel applications of NLP tools and methodologies, including uses of NLP for clinical research or operation, examples including pharmacovigilance, clinical decision support, shared decision making, computable phenotyping, predictive modeling, risk prediction, and social media mining.
Organizers
Chairs:
- Hua Xu, PhD, The University of Texas Health Science Center at Houston, US
- V.G.Vinod Vydiswaran, PhD, University of Michigan, US
Steering committee:
- Wendy Chapman, PhD, University of Utah, US
- Dina Demner Fushman, MD, PhD, National Library of Medicine, US
- Stephane Meystre, MD, PhD, Medical University of South Carolina, US
- Hongfang Liu, PhD, Mayo Clinic, US
- Guergana Savova, PhD, Harvard University, US
- Buzhou Tang, PhD, Harbin Institute of Technology, China
- Ozlem Uzuner, PhD, George Mason University, US
- Karin Verspoor, PhD, University of Melbourne, Australia
- Pierre Zweigenbaum, PhD, France
Publication committee:
- Yaoyun Zhang, PhD, The University of Texas Health Science Center at Houston, US
- Yanshan Wang, PhD, Mayo Clinic, US
Program committee:
- Samir Abdelrahman, University of Utah
- John Aberdeen, MITRE
- Eiji Aramaki, NAIST
- Tanmay Basu, Ramakrishna Mission, Vivekananda University
- Kevin Bretonnel Cohen, University of Colorado
- Wendy Chapman, University of Utah
- Cheryl Clark, MITRE
- Trevor Cohen, University of Texas Health Science Center at Houston
- Mike Conway, University of Utah
- Hercules Dalianis, DSV-Stockholm University
- Dina Demner, U.S. National Library of Medicine, NIH
- Scott Duvall, Veterans Affairs
- Carsten Eickhoff, Brown University
- Jung-Wei Fan, Kaiser Permanente
- Cyril Grouin, LIMSI, CNRS, Université Paris-Saclay
- Lynette Hirschman, MITRE
- Chun-Nan Hsu, University of California San Diego
- Zhengxing Huang, Zhejiang University
- Stephen Johnson, Weill Cornell Medicine
- Yoshinobu Kano, Shizuoka University
- Rohit Kate, Univeristy of Wisconsin-Milwaukee
- Dimitrios Kokkinakis, University of Gothenburg
- Alberto Lavelli, FBK-irst
- Jiao Li, Chinese Academy of Medical Sciences
- Feifan Liu, University of Massachusetts
- Zhiyong Lu, NCBI
- Eneida Mendonca, University of Wisconsin-Madisonordinary
- Stéphane Meystre, Medical University of South Carolina
- Tomoko Ohkuma, Fuji xerox co., Ltd.
- Jon Patrick, Health Language Analytics Global
- Tong Ruan, ECUST
- Stefan Schulz, Institute of Medical Informatics, Statistics, and Documentation, Medical University of Graz
- Isabel Segura Bedmar, Universidad Carlos III de Madrid
- Min Song, Yonsei University
- Buzhou Tang, Shenzhen Graduate School, Harbin Institute of Technology
- Ozlem Uzuner, George Mason University
- Karin Verspoor, The University of Melbourne
- V.G.Vinod Vydiswaran, University of Michigan
- Wei Wang, University of California, Los Angeles
- Yanshan Wang, Mayo Clinic
- Chunhua Weng, Columbia University
- Yonghui Wu, University of Florida
- Hua Xu, University of Texas Health Science Center at Houston
- Zhihao Yang, Dalian university of technology
- Kunli Zhang, Zhengzhou University
- Yaoyun Zhang, University of Texas Health Science Center at Houston
- Deyu Zhou, Southeast University
- Li Zhou, Brigham and Women's Hospital
- Vivienne Zhu, The Medical University of South Carolina
More to be updated...
Important dates
- Deadline for all submissions: April 12th, 2018
- Notification of decisions: April 27th, 2018
- Camera-ready submissions: May 14th, 2018
- Workshop date: June 7th, 2018
Submission and Review
Authors may submit regular papers (4-10 pages, including references) and abstracts (2 pages, including references) in the format used for the ICHI submissions. Developers of NLP tools may submit a demo paper (up to 4 pages, including references) describing the tool, along with a link to a web demonstration (interactive website or a demonstration video). As with ICHI, the workshop will follow single-blind reviewing, therefore submissions should have the names and affiliations of authors listed on the paper. Papers must adhere to the IEEE Proceedings Format available for LaTeX and Word, and submitted as a single PDF file. Submissions will be handled electronically through EasyChair (https://easychair.org/conferences/?conf=healthnlp2018). For more information, please read the detailed submission instructions (link here).
All submissions will undergo a peer review by at least two domain experts from the program committee. The selection of papers and posters will be primarily based on the potential to generate discussion and impact future research directions. Accepted papers will be included in the ICHI 2018 Proceedings and IEEE Digital Library. Plans are underway to also invite authors of selected papers to publish an extended version of their work in a special issue of BMC Medical Informatics and Decision Making journal.

 TOP
TOP